Your DNA sequence is largely fixed from conception. But the instructions encoded in that sequence — which genes are expressed in which tissues, at what levels, and at what times — are regulated by a layer of chemical modifications called the epigenome. Unlike the genome, the epigenome is dynamic, responsive to environment, and critically, modifiable by the choices you make every day. This is the molecular basis of epigenetics, and it has profound implications for cancer.
What Epigenetics Means
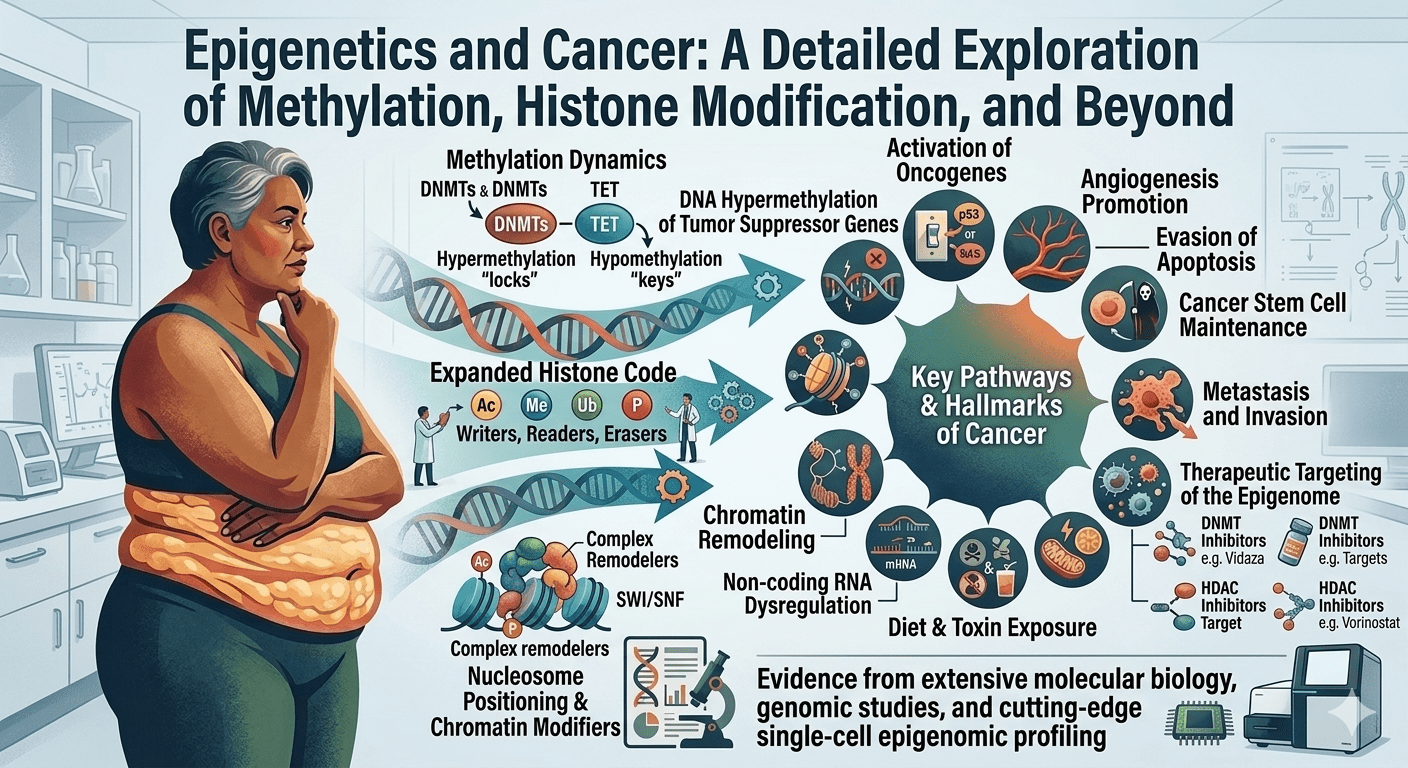
Epigenetic regulation occurs through three primary mechanisms: DNA methylation (covalent addition of methyl groups to cytosine residues, typically at CpG dinucleotides), histone modification (acetylation, methylation, phosphorylation, and ubiquitination of histone proteins around which DNA is wound), and non-coding RNA regulation (including microRNAs and long non-coding RNAs that modulate gene expression post-transcriptionally) [1].
These modifications determine whether a gene is accessible for transcription or silenced. In cancer, epigenetic dysregulation is universal: hypermethylation of tumour suppressor gene promoters silences genes like CDKN2A, BRCA1, MLH1, and VHL [4]. Global DNA hypomethylation causes chromosomal instability and reactivation of oncogenes [4]. Histone deacetylation (mediated by HDACs) compacts chromatin and silences genes involved in differentiation and apoptosis [5].
Diet as an Epigenetic Regulator
A 2018 paper in Nature Communications directly addressed the molecular interface between diet and the epigenome, characterising how dietary metabolites serve as substrates or cofactors for epigenetic enzymes [2]. Acetyl-CoA (derived from glucose and acetate metabolism) is the acetyl donor for histone acetyltransferases — meaning that dietary carbohydrate and fat intake directly influences the pool of acetyl groups available for histone modification. S-adenosylmethionine (SAM), the universal methyl donor for DNA methyltransferases, is synthesised from methionine and requires folate, B12, and choline as cofactors. Dietary deficiency in these methyl donors directly impairs DNA methylation fidelity [2].
Vitamins, Polyphenols, and Epigenetic Drugs
A 2020 critical review in Critical Reviews in Food Science and Nutrition evaluated the evidence for vitamins and dietary compounds as epidrugs — agents that modulate epigenetic marks in ways that may prevent or reverse cancer-associated epigenetic changes [3]. The review found that folate (which supports DNA methylation), retinoids (which regulate chromatin accessibility through RAR nuclear receptors), and several polyphenols — sulforaphane as an HDAC inhibitor, resveratrol as a sirtuin activator, and EGCG as a DNMT inhibitor — show biologically plausible epigenetic anti-cancer mechanisms. The authors cautioned that clinical translation requires careful attention to dose, bioavailability, and the context-dependency of epigenetic effects [3].
The Epigenetic Clock and Biological Age
One of the most striking developments in epigenetics is the discovery of DNA methylation-based biological age clocks. The Horvath clock, introduced in 2013, uses methylation patterns at 353 CpG sites to estimate biological age across multiple human tissues and cell types [6]. The Levine PhenoAge model extended this framework by integrating clinical biomarkers to produce a multi-tissue predictor that correlates more strongly with mortality and disease risk than chronological age alone [7]. These clocks measure the degree of epigenetic drift from a youthful baseline state and provide a measure of biological age that can diverge substantially from chronological age. Accelerated epigenetic ageing — driven by chronic stress, poor nutrition, sleep disruption, smoking, and physical inactivity — is associated with substantially elevated cancer risk [8]. Conversely, interventions including exercise, dietary improvement, and psychosocial stress reduction slow epigenetic ageing [8].
Epigenetic Inheritance: Passing It On
Epigenetic marks are not entirely reset between generations. Epidemiological studies of famines, toxin exposures, and maternal environments show that epigenetic adaptations can be transmitted across one to three generations — a phenomenon called transgenerational epigenetic inheritance [9]. This has implications for cancer risk: adverse exposures in parents or grandparents may influence cancer susceptibility in offspring through epigenetic mechanisms, independently of DNA sequence variants [9].
Clinical Epigenetics
Epigenetic targets are now validated drug targets. DNMT inhibitors (azacitidine, decitabine) and HDAC inhibitors (vorinostat, romidepsin) are approved cancer treatments, primarily for haematological malignancies [10]. EZH2 inhibitors (tazemetostat) target histone H3K27 methyltransferase activity in certain lymphomas [11]. The epigenome is not merely a passive record of your life — it is an active, targetable system.
References
- Feinberg AP (2018) The key role of epigenetics in human disease prevention and mitigation. N Engl J Med 378:1323–1334. doi:10.1056/NEJMra1402513.
- Zhang T and Kutateladze TG (2018) Diet and the epigenome. Nat Commun 9:3375. doi:10.1038/s41467-018-05778-1.
- Nur A, Wahid MI, Mutalip MHHM, Chin SF and Jamal R (2020) Vitamins as epidrugs: an update on epigenetic efficacy of vitamins. Crit Rev Food Sci Nutr 61:3843–3857. doi:10.1080/10408398.2020.1712674.
- Baylin SB and Jones PA (2011) A decade of exploring the cancer epigenome: biological and translational implications. Nat Rev Cancer 11:726–738. doi:10.1038/nrc3130.
- Kanwal R and Gupta S (2012) Epigenetic modifications in cancer. Clin Genet 81:303–311. doi:10.1111/j.1399-0004.2011.01809.x.
- Horvath S (2013) DNA methylation age of human tissues and cell types. Genome Biol 14:R115. doi:10.1186/gb-2013-14-10-r115.
- Levine ME, Lu AT, Quach A, Chen BH, Assimes TL, Bandinelli S, Hou L, Baccarelli AA, Stewart JD, Li Y, Whitsel EA, Wilson JG, Reiner AP, Aviv A, Liang K, Liu D, Ferrucci L and Horvath S (2018) An epigenetic biomarker of aging for lifespan and healthspan. Aging 10:573–591. doi:10.18632/aging.101414.
- Alegría-Torres JA, Baccarelli A and Bollati V (2011) Epigenetics and lifestyle. Epigenomics 3:267–277. doi:10.2217/epi.11.22.
- Skinner MK, Manikkam M and Guerrero-Bosagna C (2010) Epigenetic transgenerational actions of environmental factors in disease etiology. Trends Endocrinol Metab 21:214–222. doi:10.1016/j.tem.2009.12.007.
- Dawson MA and Kouzarides T (2012) Cancer epigenetics: from mechanism to therapy. Cell 150:12–27. doi:10.1016/j.cell.2012.06.013.
- Morera L, Lübbert M and Jung M (2016) Targeting histone methyltransferases and demethylases in clinical trials for cancer therapy. Clin Epigenetics 8:57. doi:10.1186/s13148-016-0223-4.